Cloning-Growth and Plasmid Extraction by Alkaline Lysis Method
Biology | Biochemistry | Genetics | Microbiology





2.5M+
Active Users Worldwide
80%
Improved Learning Retention
60%
Reduction in Laboratory Costs
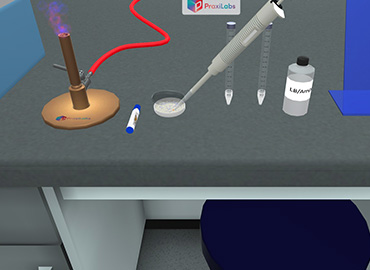
Isolation of plasmid DNA by alkaline lysis method.
Alkaline Lysis Method for Plasmid DNA Isolation
To apply the sterile technique while growing the bacterial culture.
Stages of Plasmid Extraction by Alkaline Lysis Method:
1- Resuspension of Cells:
2- Cell lysis:
3- Neutralization of the reaction and clearing of the lysate:
4- Precipitation and De-Salting:




